
Cluster analysis (continued)
- Complete linkage (furthest neighbor) clustering
- Identical to single linkage clustering except that the
distance between entities is defined as the point of maximum
distance
- e.g., distances:
| | 1 | 2 | 3 | 4 | 5 | 6 |
| 1 | - | 3.16 | 4.47 | 15.16 | 11.40 | 12.32 |
| 2 | | - | 7.07 | 12.04 | 8.94 | 9.85 |
- All distances of entities after quadrats 1 and 2 are joined:
| | 1,2 | 3 | 4 | 5 | 6 |
| 1,2 | - | 7.07 | 15.16 | 11.40 | 12.32 |
| i.e., | d(1,3)
= 4.47 |
| | d(2,3)
= 7.07; |
| thus, | d[(1,2),3] = 7.07 |
- w/ single linkage clustering, dist = minimum distance -
-> d[(1,2),3] = 4.47
- Decision rule is still based on smallest distance, but
distances are calculated differently
- Characteristics of complete linkage clustering:
- "space-dilating"--as a cluster grows it tends to become
more dissimilar to others --> non-chaining
- group structure is ignored; as w/ single linkage
clustering, comparisons are based on indiv. quadrats
- results often similar to "minimum-variance" clustering
- Centroid clustering
- Distance between 2 clusters is defined as the euclidean
distance between their centroids
- Two groups are joined if the distance between their
centroids is the smallest of all possible "choices"
- e.g., distance between groups:
- To calculate centroid:
| Quadrat | Species A | Species
B |
| 1 | 15 | 9 |
| 2 | 12 | 8 |
| 3 | 17 | 13 |
| 4 | 0 | 7 |
| 5 | 8 | 0 |
| 6 | 3 | 12 |
- First step is identical to single linkage clustering, since
groups are single quadrats. After quadrats 1 and 2 are
joined, centroid(1,2) = [(15+12)/2, (9+8)/2] = (13.5,8.5).
- centroid (1,2,3) = [(15+12+17)/3, (9+8+13)/3] =
(14.333,10)
- Then euclidean distance is calculated not between
nearest quadrats in group (single linkage) and not
between furthest quadrats in group (complete linkage),
but between centroids of groups
- Thus, group structure is used in determining between-
group similarities
- A disadvantage of centroid clustering is the potential for
reversals
- After a fusion, the next fusion occurs at a less
dissimilar point (i.e., closer distance)
- e.g., consider these 3 quadrats w/ 2 species:
| Quadrat | Species A | Species B |
| 1 | 26 | 10 |
| 2 | 34 | 6 |
| 3 | 34 | 15 |
- d(1,2) =
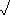 [(26-34)2 + (10-6)2] = 8.944
[(26-34)2 + (10-6)2] = 8.944
- d(1,3) =
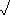 [(26-34)2 + (10-15)2] = 9.434
[(26-34)2 + (10-15)2] = 9.434
- d(2,3) =
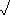 [(34-34)2 + (6-15)2] = 9.000
[(34-34)2 + (6-15)2] = 9.000
- Quadrats 1 and 2 are joined --> centroid =
[(26+34)/2,(10+6)/2] = (30,8)
- d[(1,2),3] =
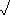 [(30-34)2 + (8-15)2] = 9.062
[(30-34)2 + (8-15)2] = 9.062
- Thus, the second fusion occurs at a smaller
distance than the first fusion (i.e., this
indicates these entities are more similar than
those joined by the first
fusion):
- Centroid clustering incorporates information about the group
when joining groups (vs. single linkage and complete linkage
clustering, which do not)
- However, reversals create interpretational difficulties, and
this has discouraged widespread use of clustering techniques
which have potential to show reversals
- Comparison studies have shown that single linkage and
centroid clustering behave similarly
- Minimum-variance clustering (Ward's method) (syn. Orloci's method
in ecological literature)
- Concept: we can measure the sum of the distances2 of
the
members of a group from the group centroid as an
indicator of group heterogeneity or dispersion
- Distance (similarity) measure: euclidean distance
- Fusion rule: groups are joined only if the increase in
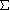 d2
is less for that pair of groups than for any
other pair
d2
is less for that pair of groups than for any
other pair
- Ward's method lends itself to a measure of "classification
efficiency":
- SStotal =
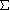 d2
of all quadrats from centroid
d2
of all quadrats from centroid
- At any point in the analysis, SS can be calculated for
each group (i.e., within-group heterogeneity or
dispersion)
- Thus, a percentage can be calculated which
indicates the proportion of total variability
explained by each group:
SSgroup/SStotal
- Characteristics of Ward's method
- Minimizes dispersion within groups
- Like complete linkage clustering, it favors the
formation of small clusters of approximately equal size
- Incorporates information about groups, not merely about
individual quadrats
- Computationally complex and time-consuming compared to
other methods we've discussed
- Widely applied in ecology, especially recently (since
computers have overcome problems w/ computational
complexity)
Previous
lectureNext lecture
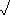 [(26-34)2 + (10-6)2] = 8.944
[(26-34)2 + (10-6)2] = 8.944
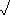 [(26-34)2 + (10-15)2] = 9.434
[(26-34)2 + (10-15)2] = 9.434
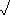 [(34-34)2 + (6-15)2] = 9.000
[(34-34)2 + (6-15)2] = 9.000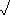 [(30-34)2 + (8-15)2] = 9.062
[(30-34)2 + (8-15)2] = 9.062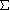 d2
is less for that pair of groups than for any
other pair
d2
is less for that pair of groups than for any
other pair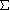 d2
of all quadrats from centroid
d2
of all quadrats from centroid